SUMMARY
Deep learning and uncertainty estimation (via Monte Carlo dropout and cross-validation) to classify digital pathology images by confidence, improving reliability in cancer diagnostics and supporting effective clinical decision-making
The Unmet Need: Framework for enhancing reliability of deep learning model predictions in clinical cancer diagnostics
- The field of digital histopathology has rapidly adopted deep learning to support diagnostic efforts, yet the inherent variability in whole-slide imaging and staining processes has exposed significant challenges. Many current methods lack mechanisms to gauge prediction reliability, leaving practitioners uncertain about the trustworthiness of automated diagnoses. This scarcity of robust uncertainty measures is especially problematic when models encounter images that differ from the training data, undermining confidence in clinical decision-making and risking potential misdiagnoses in diverse patient populations.
-
Existing approaches often focus narrowly on limited tissue sections and struggle to account for broader variations seen in whole slides or across patients. Moreover, these models frequently face issues such as overfitting to controlled datasets and data leakage during threshold calibration, which hinder their performance on external or synthetic datasets. This inadequacy raises substantial concerns about the consistency and safety of deploying deep learning systems in clinical settings, where accurate identification of uncertain predictions is crucial for timely human intervention and optimal patient care.
The proposed solution: A combination of a training schema and thresholding algorithm used to quantity the amount of uncertainty in deep learning model’s predictions
- This deep learning technology for digital histopathology leverages an Xception-based convolutional neural network enhanced with Monte Carlo dropout and nested cross-validation to quantify uncertainty. It processes whole-slide images with robust preprocessing, such as stain normalization, and computes performance metrics including AUROC and activation maps. The system refines uncertainty measures by classifying predictions as low or high confidence at a slide- or patient-level, ensuring the use of non-leaked training data for threshold definition. This approach allows reliable predictions even when encountering out-of-distribution data, including external datasets and synthetic images near decision boundaries.
-
What sets this technology apart is its comprehensive evaluation framework and advanced uncertainty quantification. By integrating Monte Carlo dropout to generate a predictive distribution and deriving Bayesian-inspired uncertainty metrics, it provides a rigorous and transparent method for assessing prediction reliability. Extensive multi-institutional validation and synthetic testing confirm its robustness, particularly as it addresses domain shifts that typically complicate digital pathology. This differentiated performance supports better clinical decision-making, reducing reliance on manual reviews while highlighting clear morphological correlations with high-confidence predictions.
FIGURE
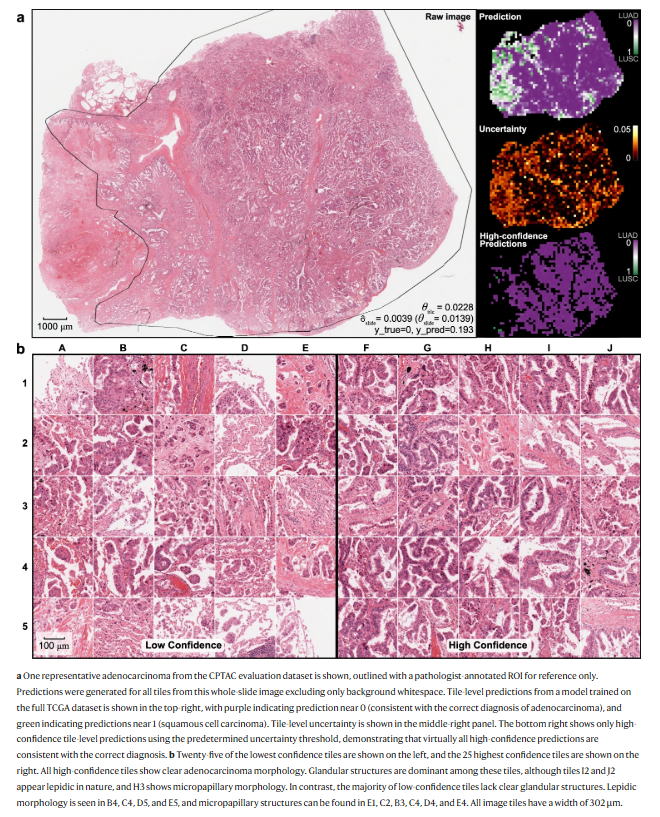
ADVANTAGES
ADVANTAGES
- Enhances diagnostic reliability by quantifying uncertainty at the slide or patient level
-
Addresses domain shift challenges, ensuring robust performance across various datasets and institutions
-
Prevents data leakage by establishing thresholds exclusively on training data through nested cross-validation
-
Integrates advanced techniques like Monte Carlo dropout to produce Bayesian uncertainty estimates
-
Improves clinical decision-making by flagging low-confidence predictions for manual review or alternative treatment paths
APPLICATIONS
- Digital pathology diagnostic support
- Clinical cancer risk prediction
- Automated histopathology slide review
- Uncertainty-driven treatment decisions
- Deep learning pathology analytics
PUBLICATIONS
February 28, 2024
Proof of concept
Patent Pending
Licensing,Co-development
Alexander Pearson